import numpy as np
from pysal.lib import weights
import pandas as pd
import shapely as sp
import contextily as ctx
import geopandas as gpd
import networkx as nx
import matplotlib.pyplot as plt
import esda
from splot.esda import plot_moran, lisa_cluster
from pysal.model import spregSpatial Statistics
Spatial statistics
Outline for this part: - Graphs & spatial weight matrices - Spatial autocorrelation - Standard geographic regression models
constituencies_shapes = gpd.read_file("./data/constituencies_shape/circonscriptions_legislatives_030522.shp")
constituencies_shapes = constituencies_shapes[~constituencies_shapes["id_circo"].str.startswith("97")]
constituencies_shapes.explore()constituencies_x = pd.read_csv("./data/constituencies_x.csv")
constituencies_x["id_circo"] = constituencies_x["id_circo"].astype(str).str[0:2] + constituencies_x["id_circo"].astype(str).str[3:5]
constituencies_y = pd.read_csv("./data/constituencies_y.csv")
constituencies = constituencies_shapes \
.merge(constituencies_x, on = "id_circo", how = "left") \
.merge(constituencies_y, on = "id_circo", how = "left") constituencies| id_circo | dep | libelle | geometry | Nom de la circonscription | mean_age | unemployement_rate | D1_diff | D9_diff | rpt_D9_D1_diff | vote_share_rn | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 3803 | 38 | Isère - 3e circonscription | MULTIPOLYGON (((5.70136 45.18796, 5.70136 45.1... | Isère - 3e circonscription | 38.1 | 7.2 | 10530 | 36370 | 3.5 | 20.41 |
| 1 | 3801 | 38 | Isère - 1re circonscription | MULTIPOLYGON (((5.74302 45.16638, 5.74259 45.1... | Isère - 1re circonscription | 39.8 | 5.4 | 12080 | 51150 | 4.2 | 14.39 |
| 2 | 3810 | 38 | Isère - 10e circonscription | MULTIPOLYGON (((5.67138 45.47904, 5.66725 45.4... | Isère - 10e circonscription | 37.8 | 6.1 | 12240 | 36130 | 3 | 38.43 |
| 3 | 3804 | 38 | Isère - 4e circonscription | MULTIPOLYGON (((5.80147 44.70678, 5.77463 44.6... | Isère - 4e circonscription | 41.9 | 4.5 | 13160 | 39110 | 3 | 28.41 |
| 4 | 3802 | 38 | Isère - 2e circonscription | MULTIPOLYGON (((5.87334 45.09739, 5.88031 45.0... | Isère - 2e circonscription | 39.0 | 5.8 | 11760 | 37610 | 3.2 | 27.67 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 534 | 0502 | 05 | Hautes-Alpes - 2e circonscription | MULTIPOLYGON (((6.23277 44.46288, 6.24026 44.4... | Hautes-Alpes - 2e circonscription | 44.6 | 3.8 | 11980 | 35190 | 2.9 | 29.44 |
| 535 | 0401 | 04 | Alpes-de-Haute-Provence - 1re circonscription | MULTIPOLYGON (((6.84988 43.9141, 6.83595 43.91... | Alpes de Haute-Provence - 1re circonscription | 45.9 | 6.0 | 11310 | 34120 | 3 | 38.50 |
| 536 | 4204 | 42 | Loire - 4e circonscription | MULTIPOLYGON (((4.54255 45.24218, 4.53626 45.2... | Loire - 4e circonscription | 41.9 | 4.9 | 12140 | 34470 | 2.8 | 38.62 |
| 537 | 0703 | 07 | Ardèche - 3e circonscription | POLYGON ((4.05709 44.36415, 4.05709 44.36415, ... | Ardèche - 3e circonscription | 46.7 | 7.3 | 10880 | 33580 | 3.1 | 32.02 |
| 538 | 0501 | 05 | Hautes-Alpes - 1re circonscription | MULTIPOLYGON (((6.26146 44.5461, 6.26146 44.54... | Hautes-Alpes - 1re circonscription | 44.5 | 5.4 | 11770 | 35350 | 3 | 32.56 |
539 rows × 11 columns
constituencies.set_index('id_circo')| dep | libelle | geometry | Nom de la circonscription | mean_age | unemployement_rate | D1_diff | D9_diff | rpt_D9_D1_diff | vote_share_rn | |
|---|---|---|---|---|---|---|---|---|---|---|
| id_circo | ||||||||||
| 3803 | 38 | Isère - 3e circonscription | MULTIPOLYGON (((5.70136 45.18796, 5.70136 45.1... | Isère - 3e circonscription | 38.1 | 7.2 | 10530 | 36370 | 3.5 | 20.41 |
| 3801 | 38 | Isère - 1re circonscription | MULTIPOLYGON (((5.74302 45.16638, 5.74259 45.1... | Isère - 1re circonscription | 39.8 | 5.4 | 12080 | 51150 | 4.2 | 14.39 |
| 3810 | 38 | Isère - 10e circonscription | MULTIPOLYGON (((5.67138 45.47904, 5.66725 45.4... | Isère - 10e circonscription | 37.8 | 6.1 | 12240 | 36130 | 3 | 38.43 |
| 3804 | 38 | Isère - 4e circonscription | MULTIPOLYGON (((5.80147 44.70678, 5.77463 44.6... | Isère - 4e circonscription | 41.9 | 4.5 | 13160 | 39110 | 3 | 28.41 |
| 3802 | 38 | Isère - 2e circonscription | MULTIPOLYGON (((5.87334 45.09739, 5.88031 45.0... | Isère - 2e circonscription | 39.0 | 5.8 | 11760 | 37610 | 3.2 | 27.67 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 0502 | 05 | Hautes-Alpes - 2e circonscription | MULTIPOLYGON (((6.23277 44.46288, 6.24026 44.4... | Hautes-Alpes - 2e circonscription | 44.6 | 3.8 | 11980 | 35190 | 2.9 | 29.44 |
| 0401 | 04 | Alpes-de-Haute-Provence - 1re circonscription | MULTIPOLYGON (((6.84988 43.9141, 6.83595 43.91... | Alpes de Haute-Provence - 1re circonscription | 45.9 | 6.0 | 11310 | 34120 | 3 | 38.50 |
| 4204 | 42 | Loire - 4e circonscription | MULTIPOLYGON (((4.54255 45.24218, 4.53626 45.2... | Loire - 4e circonscription | 41.9 | 4.9 | 12140 | 34470 | 2.8 | 38.62 |
| 0703 | 07 | Ardèche - 3e circonscription | POLYGON ((4.05709 44.36415, 4.05709 44.36415, ... | Ardèche - 3e circonscription | 46.7 | 7.3 | 10880 | 33580 | 3.1 | 32.02 |
| 0501 | 05 | Hautes-Alpes - 1re circonscription | MULTIPOLYGON (((6.26146 44.5461, 6.26146 44.54... | Hautes-Alpes - 1re circonscription | 44.5 | 5.4 | 11770 | 35350 | 3 | 32.56 |
539 rows × 10 columns
paris = constituencies.query("dep == '75'").to_crs(epsg = "2154")
paris.explore()Graphs & spatial weight matrices
Mathematical representation of geometries
One may want to encode as mathematical objects, e.g. numbers,some relationship between two geographic units:
- Are unit A and B neighbours?
- How “far” is unit A from unit B?
- How “strong” is the relationship between unit A and unit B?
Geometries are not useful for this, but graphs are.
Graphs are a data structure with nodes and a set of connection between them called edges.
In our case, nodes might be geometric objects and the relationships (or absence of) between them the edges.
Are two geometries neighbours?
One possible definition may be that A and B are neighbours if they share at least one vertex or one edge
W = weights.contiguity.Queen.from_dataframe(paris, idVariable = "id_circo")/tmp/ipykernel_26467/3174741056.py:1: FutureWarning: `idVariable` is deprecated and will be removed in future. Use `ids` instead.
W = weights.contiguity.Queen.from_dataframe(paris, idVariable = "id_circo")centroids = np.column_stack((paris.centroid.x, paris.centroid.y))
graph = W.to_networkx()
positions = dict(zip(graph.nodes, centroids))
# plot with a nice basemap
ax = paris.plot(linewidth=1, edgecolor="grey", facecolor="white")
nx.draw(graph, positions, ax=ax, node_size=5, node_color="r")
plt.show()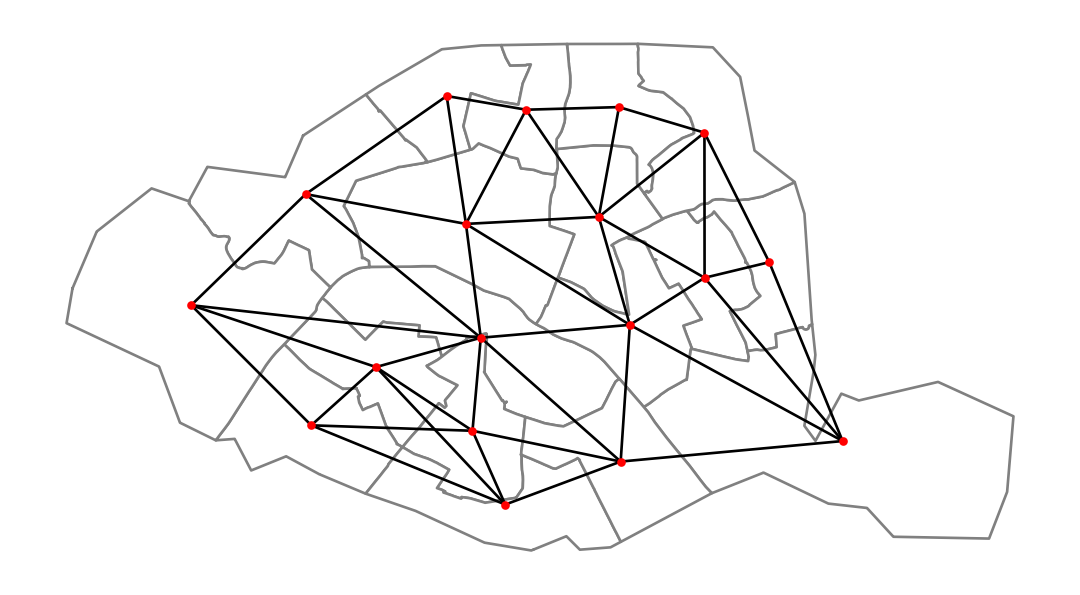
W.neighbors{'7506': ['7515', '7508', '7516', '7505', '7507'],
'7505': ['7501', '7517', '7516', '7506', '7518', '7507'],
'7502': ['7511', '7501', '7512', '7514', '7509', '7504', '7507'],
'7501': ['7502', '7504', '7503', '7505', '7518', '7507'],
'7504': ['7502', '7501', '7503', '7514'],
'7517': ['7516', '7505', '7518'],
'7512': ['7502', '7511', '7514', '7510', '7513'],
'7515': ['7508', '7516', '7506'],
'7511': ['7502', '7512', '7509', '7510', '7513'],
'7513': ['7511', '7510', '7512', '7514'],
'7508': ['7509', '7507', '7506', '7515'],
'7507': ['7502', '7501', '7505', '7508', '7509', '7506'],
'7503': ['7501', '7504', '7518'],
'7516': ['7505', '7517', '7506', '7515'],
'7518': ['7501', '7517', '7505', '7503'],
'7514': ['7502', '7504', '7513', '7512'],
'7510': ['7511', '7509', '7513', '7512'],
'7509': ['7502', '7511', '7508', '7510', '7507']}W.weights{'7506': [1.0, 1.0, 1.0, 1.0, 1.0],
'7505': [1.0, 1.0, 1.0, 1.0, 1.0, 1.0],
'7502': [1.0, 1.0, 1.0, 1.0, 1.0, 1.0, 1.0],
'7501': [1.0, 1.0, 1.0, 1.0, 1.0, 1.0],
'7504': [1.0, 1.0, 1.0, 1.0],
'7517': [1.0, 1.0, 1.0],
'7512': [1.0, 1.0, 1.0, 1.0, 1.0],
'7515': [1.0, 1.0, 1.0],
'7511': [1.0, 1.0, 1.0, 1.0, 1.0],
'7513': [1.0, 1.0, 1.0, 1.0],
'7508': [1.0, 1.0, 1.0, 1.0],
'7507': [1.0, 1.0, 1.0, 1.0, 1.0, 1.0],
'7503': [1.0, 1.0, 1.0],
'7516': [1.0, 1.0, 1.0, 1.0],
'7518': [1.0, 1.0, 1.0, 1.0],
'7514': [1.0, 1.0, 1.0, 1.0],
'7510': [1.0, 1.0, 1.0, 1.0],
'7509': [1.0, 1.0, 1.0, 1.0, 1.0]}Adjacency matrix
pd.DataFrame(*W.full()).astype(int)| 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 7506 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 1 | 0 | 1 | 0 | 0 | 0 | 0 |
| 7505 | 1 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 0 | 0 |
| 7502 | 0 | 0 | 0 | 1 | 1 | 0 | 1 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 1 |
| 7501 | 0 | 1 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 1 | 0 | 0 | 0 |
| 7504 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 |
| 7517 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 |
| 7512 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 |
| 7515 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 |
| 7511 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 |
| 7513 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 |
| 7508 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 1 |
| 7507 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
| 7503 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 |
| 7516 | 1 | 1 | 0 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| 7518 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 |
| 7514 | 0 | 0 | 1 | 0 | 1 | 0 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| 7510 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
| 7509 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 0 |
Note that we went from geometries to a matrix, which we can work with using usual linear algebra tools
W.cardinalities{'7506': 5,
'7505': 6,
'7502': 7,
'7501': 6,
'7504': 4,
'7517': 3,
'7512': 5,
'7515': 3,
'7511': 5,
'7513': 4,
'7508': 4,
'7507': 6,
'7503': 3,
'7516': 4,
'7518': 4,
'7514': 4,
'7510': 4,
'7509': 5}W.transform = "R"
pd.DataFrame(*W.full())| 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 7506 | 0.000000 | 0.200000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.20 | 0.000000 | 0.00 | 0.200000 | 0.200000 | 0.000000 | 0.200000 | 0.000000 | 0.000000 | 0.00 | 0.000000 |
| 7505 | 0.166667 | 0.000000 | 0.000000 | 0.166667 | 0.000000 | 0.166667 | 0.000000 | 0.00 | 0.000000 | 0.00 | 0.000000 | 0.166667 | 0.000000 | 0.166667 | 0.166667 | 0.000000 | 0.00 | 0.000000 |
| 7502 | 0.000000 | 0.000000 | 0.000000 | 0.142857 | 0.142857 | 0.000000 | 0.142857 | 0.00 | 0.142857 | 0.00 | 0.000000 | 0.142857 | 0.000000 | 0.000000 | 0.000000 | 0.142857 | 0.00 | 0.142857 |
| 7501 | 0.000000 | 0.166667 | 0.166667 | 0.000000 | 0.166667 | 0.000000 | 0.000000 | 0.00 | 0.000000 | 0.00 | 0.000000 | 0.166667 | 0.166667 | 0.000000 | 0.166667 | 0.000000 | 0.00 | 0.000000 |
| 7504 | 0.000000 | 0.000000 | 0.250000 | 0.250000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.000000 | 0.00 | 0.000000 | 0.000000 | 0.250000 | 0.000000 | 0.000000 | 0.250000 | 0.00 | 0.000000 |
| 7517 | 0.000000 | 0.333333 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.000000 | 0.00 | 0.000000 | 0.000000 | 0.000000 | 0.333333 | 0.333333 | 0.000000 | 0.00 | 0.000000 |
| 7512 | 0.000000 | 0.000000 | 0.200000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.200000 | 0.20 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.200000 | 0.20 | 0.000000 |
| 7515 | 0.333333 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.000000 | 0.00 | 0.333333 | 0.000000 | 0.000000 | 0.333333 | 0.000000 | 0.000000 | 0.00 | 0.000000 |
| 7511 | 0.000000 | 0.000000 | 0.200000 | 0.000000 | 0.000000 | 0.000000 | 0.200000 | 0.00 | 0.000000 | 0.20 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.20 | 0.200000 |
| 7513 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.250000 | 0.00 | 0.250000 | 0.00 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.250000 | 0.25 | 0.000000 |
| 7508 | 0.250000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.25 | 0.000000 | 0.00 | 0.000000 | 0.250000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.250000 |
| 7507 | 0.166667 | 0.166667 | 0.166667 | 0.166667 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.000000 | 0.00 | 0.166667 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.166667 |
| 7503 | 0.000000 | 0.000000 | 0.000000 | 0.333333 | 0.333333 | 0.000000 | 0.000000 | 0.00 | 0.000000 | 0.00 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.333333 | 0.000000 | 0.00 | 0.000000 |
| 7516 | 0.250000 | 0.250000 | 0.000000 | 0.000000 | 0.000000 | 0.250000 | 0.000000 | 0.25 | 0.000000 | 0.00 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.000000 |
| 7518 | 0.000000 | 0.250000 | 0.000000 | 0.250000 | 0.000000 | 0.250000 | 0.000000 | 0.00 | 0.000000 | 0.00 | 0.000000 | 0.000000 | 0.250000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.000000 |
| 7514 | 0.000000 | 0.000000 | 0.250000 | 0.000000 | 0.250000 | 0.000000 | 0.250000 | 0.00 | 0.000000 | 0.25 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.000000 |
| 7510 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.250000 | 0.00 | 0.250000 | 0.25 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.250000 |
| 7509 | 0.000000 | 0.000000 | 0.200000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.00 | 0.200000 | 0.00 | 0.200000 | 0.200000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.20 | 0.000000 |
pd.Series(W.cardinalities).plot.hist(color="k")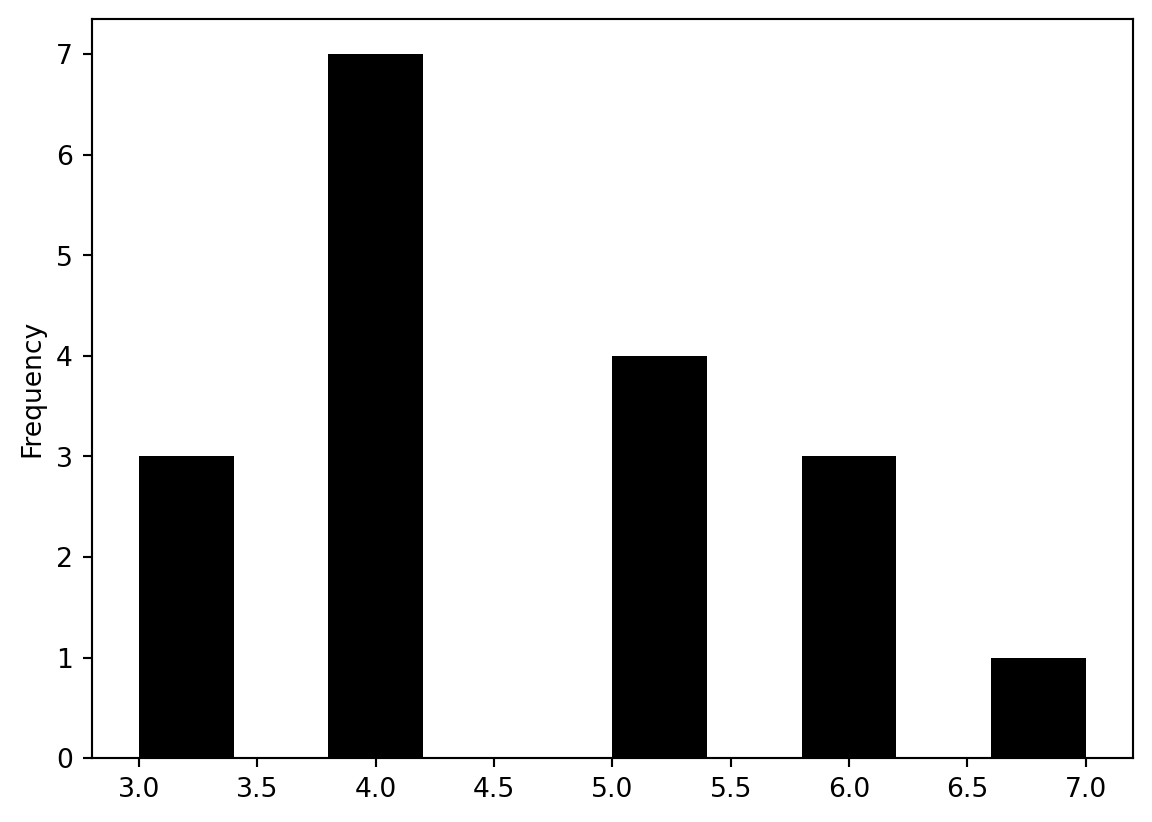
Other possible definition of weights using kernels
W = weights.distance.Kernel.from_dataframe(paris, function = "gaussian")pd.DataFrame(*W.full())| 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 52 | 0.398942 | 0.338931 | 0.000000 | 0.000000 | 0.000000 | 0.267739 | 0.000000 | 0.380092 | 0.000000 | 0.000000 | 0.241971 | 0.366320 | 0.000000 | 0.316847 | 0.000000 | 0.000000 | 0.000000 | 0.255290 |
| 53 | 0.338931 | 0.398942 | 0.291999 | 0.328731 | 0.000000 | 0.348119 | 0.000000 | 0.283863 | 0.000000 | 0.000000 | 0.000000 | 0.347477 | 0.264008 | 0.326715 | 0.332084 | 0.000000 | 0.000000 | 0.000000 |
| 55 | 0.000000 | 0.291999 | 0.398942 | 0.345396 | 0.000000 | 0.000000 | 0.350514 | 0.000000 | 0.362621 | 0.267876 | 0.000000 | 0.312283 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.292319 | 0.272336 |
| 56 | 0.000000 | 0.328731 | 0.345396 | 0.398942 | 0.298303 | 0.265964 | 0.291857 | 0.000000 | 0.249691 | 0.000000 | 0.000000 | 0.265977 | 0.332224 | 0.000000 | 0.332484 | 0.000000 | 0.000000 | 0.000000 |
| 57 | 0.000000 | 0.000000 | 0.000000 | 0.298303 | 0.398942 | 0.000000 | 0.272444 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.288869 | 0.000000 | 0.000000 | 0.302015 | 0.000000 | 0.000000 |
| 58 | 0.267739 | 0.348119 | 0.000000 | 0.265964 | 0.000000 | 0.398942 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.287942 | 0.365686 | 0.362977 | 0.000000 | 0.000000 | 0.000000 |
| 59 | 0.000000 | 0.000000 | 0.350514 | 0.291857 | 0.272444 | 0.000000 | 0.398942 | 0.000000 | 0.344688 | 0.367152 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.263290 | 0.270262 | 0.000000 |
| 60 | 0.380092 | 0.283863 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.398942 | 0.000000 | 0.000000 | 0.265076 | 0.308999 | 0.000000 | 0.317139 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
| 61 | 0.000000 | 0.000000 | 0.362621 | 0.249691 | 0.000000 | 0.000000 | 0.344688 | 0.000000 | 0.398942 | 0.300098 | 0.000000 | 0.269137 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.371497 | 0.310528 |
| 74 | 0.000000 | 0.000000 | 0.267876 | 0.000000 | 0.000000 | 0.000000 | 0.367152 | 0.000000 | 0.300098 | 0.398942 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.291013 | 0.246770 | 0.000000 |
| 289 | 0.241971 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.265076 | 0.000000 | 0.000000 | 0.398942 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
| 290 | 0.366320 | 0.347477 | 0.312283 | 0.265977 | 0.000000 | 0.000000 | 0.000000 | 0.308999 | 0.269137 | 0.000000 | 0.000000 | 0.398942 | 0.000000 | 0.250599 | 0.000000 | 0.000000 | 0.000000 | 0.324895 |
| 291 | 0.000000 | 0.264008 | 0.000000 | 0.332224 | 0.288869 | 0.287942 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.398942 | 0.000000 | 0.371598 | 0.000000 | 0.000000 | 0.000000 |
| 292 | 0.316847 | 0.326715 | 0.000000 | 0.000000 | 0.000000 | 0.365686 | 0.000000 | 0.317139 | 0.000000 | 0.000000 | 0.000000 | 0.250599 | 0.000000 | 0.398942 | 0.280075 | 0.000000 | 0.000000 | 0.000000 |
| 293 | 0.000000 | 0.332084 | 0.000000 | 0.332484 | 0.000000 | 0.362977 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.371598 | 0.280075 | 0.398942 | 0.000000 | 0.000000 | 0.000000 |
| 294 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.302015 | 0.000000 | 0.263290 | 0.000000 | 0.000000 | 0.291013 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.398942 | 0.000000 | 0.000000 |
| 295 | 0.000000 | 0.000000 | 0.292319 | 0.000000 | 0.000000 | 0.000000 | 0.270262 | 0.000000 | 0.371497 | 0.246770 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.398942 | 0.337942 |
| 296 | 0.255290 | 0.000000 | 0.272336 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.310528 | 0.000000 | 0.000000 | 0.324895 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.337942 | 0.398942 |
full_matrix, ids = W.full()
paris.assign(weight_0 = full_matrix[0]).plot("weight_0", cmap="Reds")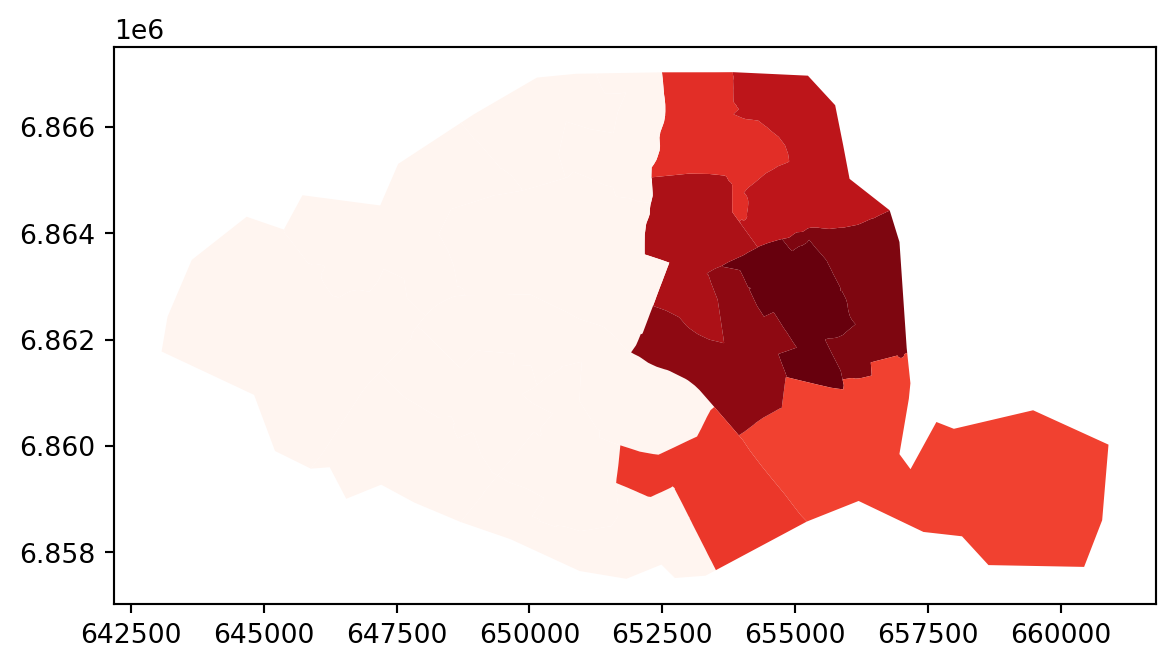
W = weights.distance.Kernel.from_dataframe(paris, function = "triangular", k=15)
full_matrix, ids = W.full()
paris.assign(weight_0 = full_matrix[0]).plot("weight_0", cmap="Reds")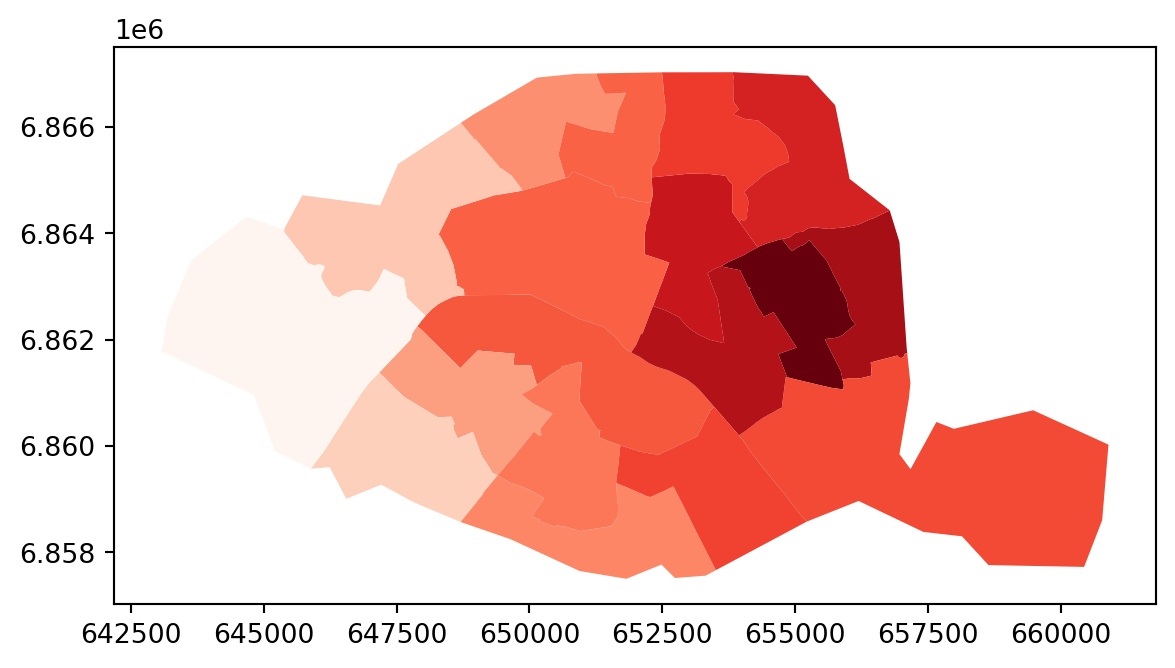
| Type | Description | Use Case |
|---|---|---|
| Queen Contiguity | Neighbors share edge OR vertex | Polygons (regions) |
| Rook Contiguity | Neighbors share edge only | Grid data |
| K-Nearest Neighbors | K closest observations | Point data |
| Distance Band | All within threshold distance | Point data |
| Kernel | Distance-weighted | Smooth spatial effects |
Practice
Take the whole
constituenciesgeoDataFrame- build the
contiguity.Queenweights matrix - What is the distribution of the number of neighbours?
- Which constituencies have the most neighbours?
- What are the neighbours of constituency
7513?
- build the
# TODO
W = weights.contiguity.Queen.from_dataframe(constituencies, idVariable = "id_circo")
pd.Series(W.cardinalities).plot.hist(color="k")
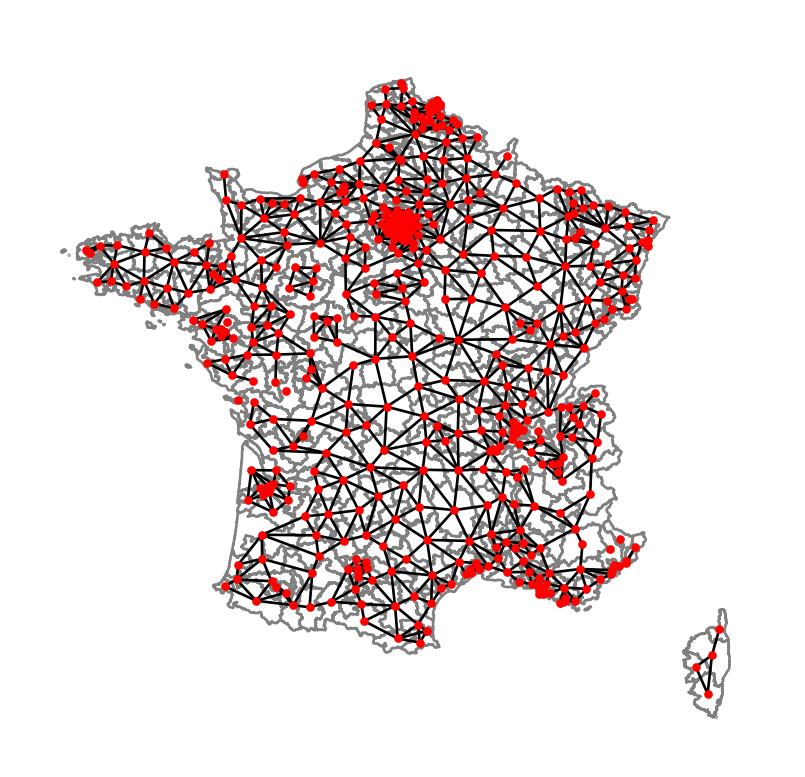
centroids = np.column_stack((constituencies.centroid.x, constituencies.centroid.y))
graph = W.to_networkx()
positions = dict(zip(graph.nodes, centroids))
# plot with a nice basemap
ax = constituencies.plot(linewidth=1, edgecolor="grey", facecolor="white")
nx.draw(graph, positions, ax=ax, node_size=5, node_color="r")
plt.show()
W.neighbors.get('7801')
{z for z in W.cardinalities if W.cardinalities[z] == 9}/tmp/ipykernel_26467/196323581.py:3: FutureWarning: `idVariable` is deprecated and will be removed in future. Use `ids` instead.
W = weights.contiguity.Queen.from_dataframe(constituencies, idVariable = "id_circo")
/opt/python/lib/python3.13/site-packages/libpysal/weights/contiguity.py:347: UserWarning: The weights matrix is not fully connected:
There are 11 disconnected components.
There are 5 islands with ids: 4405, 7902, 0602, 0605, 1701.
W.__init__(self, neighbors, ids=ids, **kw)
/tmp/ipykernel_26467/196323581.py:7: UserWarning: Geometry is in a geographic CRS. Results from 'centroid' are likely incorrect. Use 'GeoSeries.to_crs()' to re-project geometries to a projected CRS before this operation.
centroids = np.column_stack((constituencies.centroid.x, constituencies.centroid.y))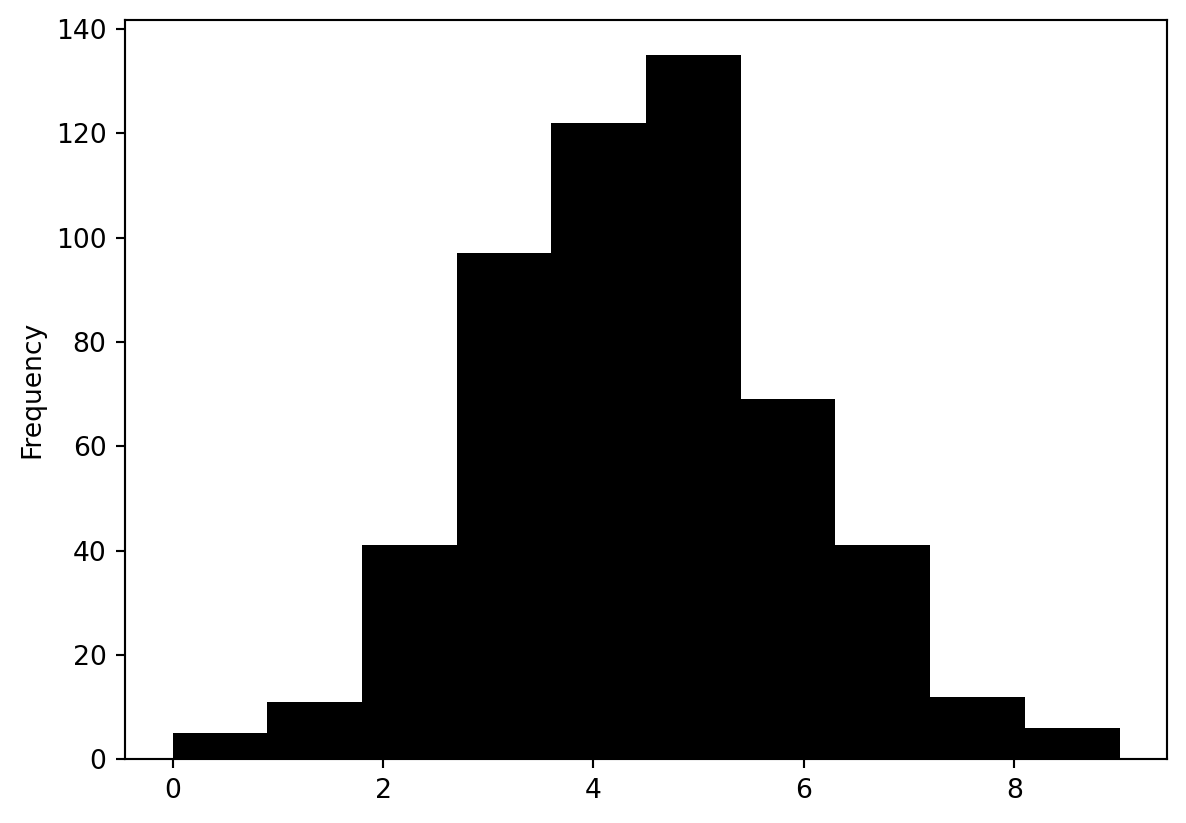
{'0205', '5704', '5802', '6201', '7602', '8004'}Spatial autocorrelation
constituencies.plot("vote_share_rn", cmap="Greys")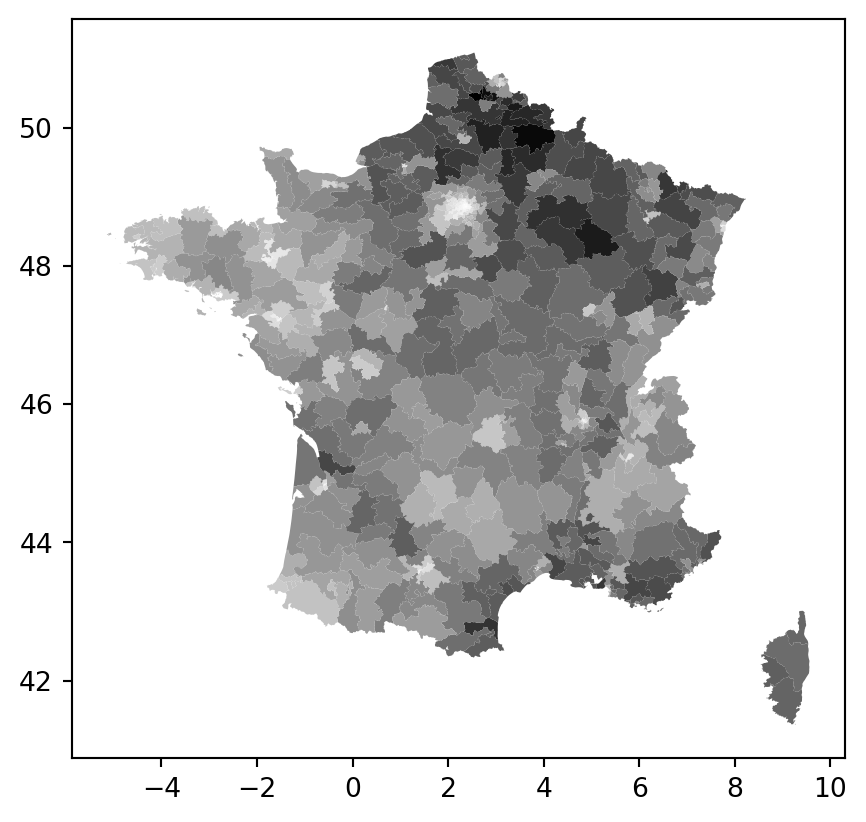
Global autocorrelation
How does the vote share of any given constituency relate to the constituencies of its neighbouring constituencies?
- are “high” vote constituencies close to “high” vote constituencies? “low” to “low”?
- or “high” to “low” and “low” to “high”?
- or pure randomness?
W = weights.contiguity.Queen.from_dataframe(constituencies, ids = "id_circo")
W.transform = "R" # row standardization: rows sum to 1('WARNING: ', '4405', ' is an island (no neighbors)')
('WARNING: ', '7902', ' is an island (no neighbors)')
('WARNING: ', '0602', ' is an island (no neighbors)')
('WARNING: ', '0605', ' is an island (no neighbors)')
('WARNING: ', '1701', ' is an island (no neighbors)')/opt/python/lib/python3.13/site-packages/libpysal/weights/contiguity.py:347: UserWarning: The weights matrix is not fully connected:
There are 11 disconnected components.
There are 5 islands with ids: 4405, 7902, 0602, 0605, 1701.
W.__init__(self, neighbors, ids=ids, **kw)Spatial lag operator (analogous to time lag operator in time series):
\(X_{lag} = WX\)
\(X_{lag} = \sum_{i} w_i x_i\)
If W is row normalized, this is a weighted averages of neighbor values using the spatial weights
We can now reframe the spatial autocorrelation as:
\(Corr(X, X_{lag})\)
constituencies["lagged_vote_share_rn"] = weights.spatial_lag.lag_spatial(W, constituencies["vote_share_rn"])constituencies[["vote_share_rn", "lagged_vote_share_rn"]]| vote_share_rn | lagged_vote_share_rn | |
|---|---|---|
| 0 | 20.41 | 25.694000 |
| 1 | 14.39 | 25.033333 |
| 2 | 38.43 | 37.555000 |
| 3 | 28.41 | 28.304000 |
| 4 | 27.67 | 22.557500 |
| ... | ... | ... |
| 534 | 29.44 | 34.086667 |
| 535 | 38.50 | 39.245000 |
| 536 | 38.62 | 35.733750 |
| 537 | 32.02 | 36.588000 |
| 538 | 32.56 | 30.520000 |
539 rows × 2 columns
constituencies.plot("lagged_vote_share_rn", cmap="Greys")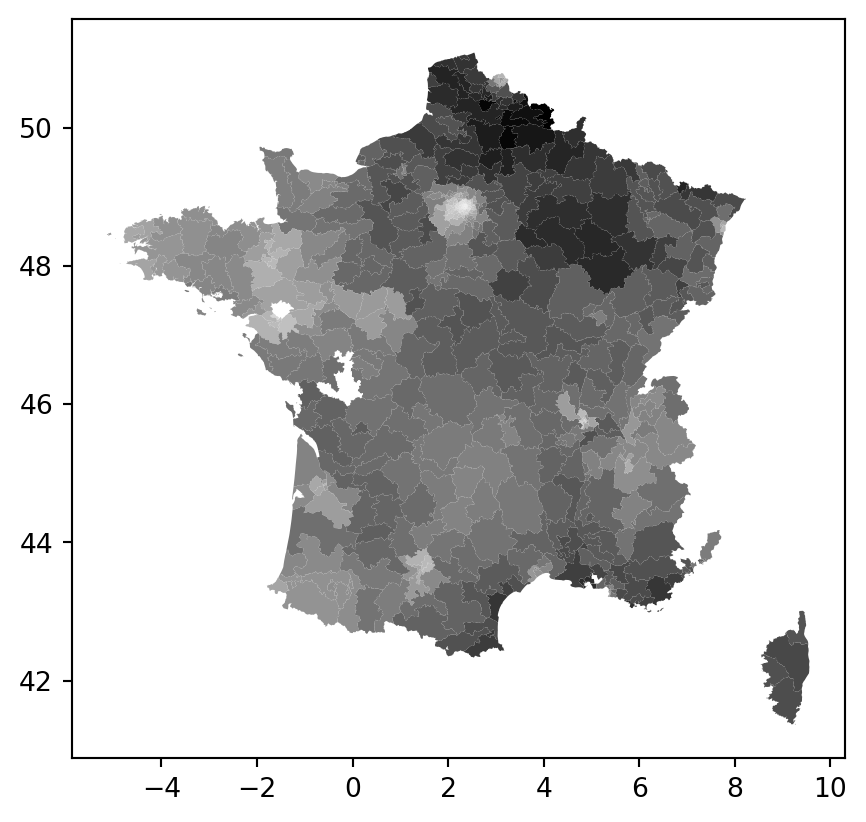
constituencies["vote_share_rn_standardized"] = constituencies["vote_share_rn"] - constituencies["vote_share_rn"].mean()
constituencies["lagged_vote_share_rn_standardized"] = weights.lag_spatial(W, constituencies["vote_share_rn_standardized"])
fig, ax = plt.subplots(figsize=(9, 9))
ax.scatter(constituencies["vote_share_rn_standardized"], constituencies["lagged_vote_share_rn_standardized"])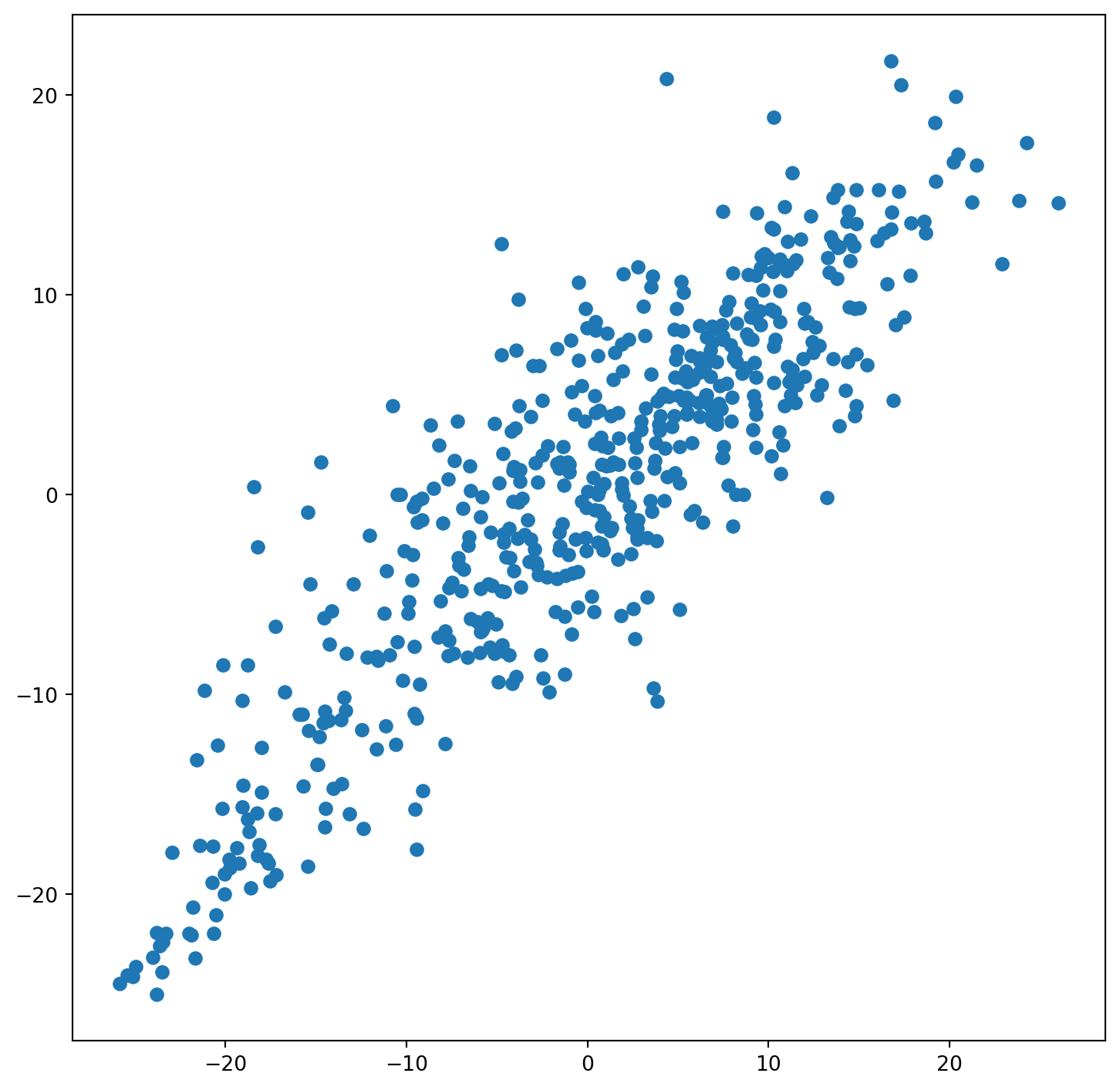
constituencies[["vote_share_rn", "lagged_vote_share_rn"]].corr()| vote_share_rn | lagged_vote_share_rn | |
|---|---|---|
| vote_share_rn | 1.00000 | 0.83766 |
| lagged_vote_share_rn | 0.83766 | 1.00000 |
Similar concept with good statistical properties: Moran’s I
\(I = \frac{n \sum_i \sum_j w_{ij}(Y_i - \bar Y)(Y_j - \bar Y)} {(\sum_{i \neq j} w_{ij}) \sum_i (Y_i - \bar Y)^2} = \frac{n}{\sum_{i \neq j} w_{ij}} \frac{\sum_i \sum_j w_{ij} z_i z_j}{\sum_i z_i^2}\)
moran = esda.moran.Moran(constituencies["vote_share_rn"], W)
moran.Inp.float64(0.7794561095576024)moran.p_simnp.float64(0.001)plot_moran(moran)(<Figure size 960x384 with 2 Axes>,
array([<Axes: title={'center': 'Reference Distribution'}, xlabel='Moran I: 0.78', ylabel='Density'>,
<Axes: title={'center': 'Moran Scatterplot (0.78)'}, xlabel='Attribute', ylabel='Spatial Lag'>],
dtype=object))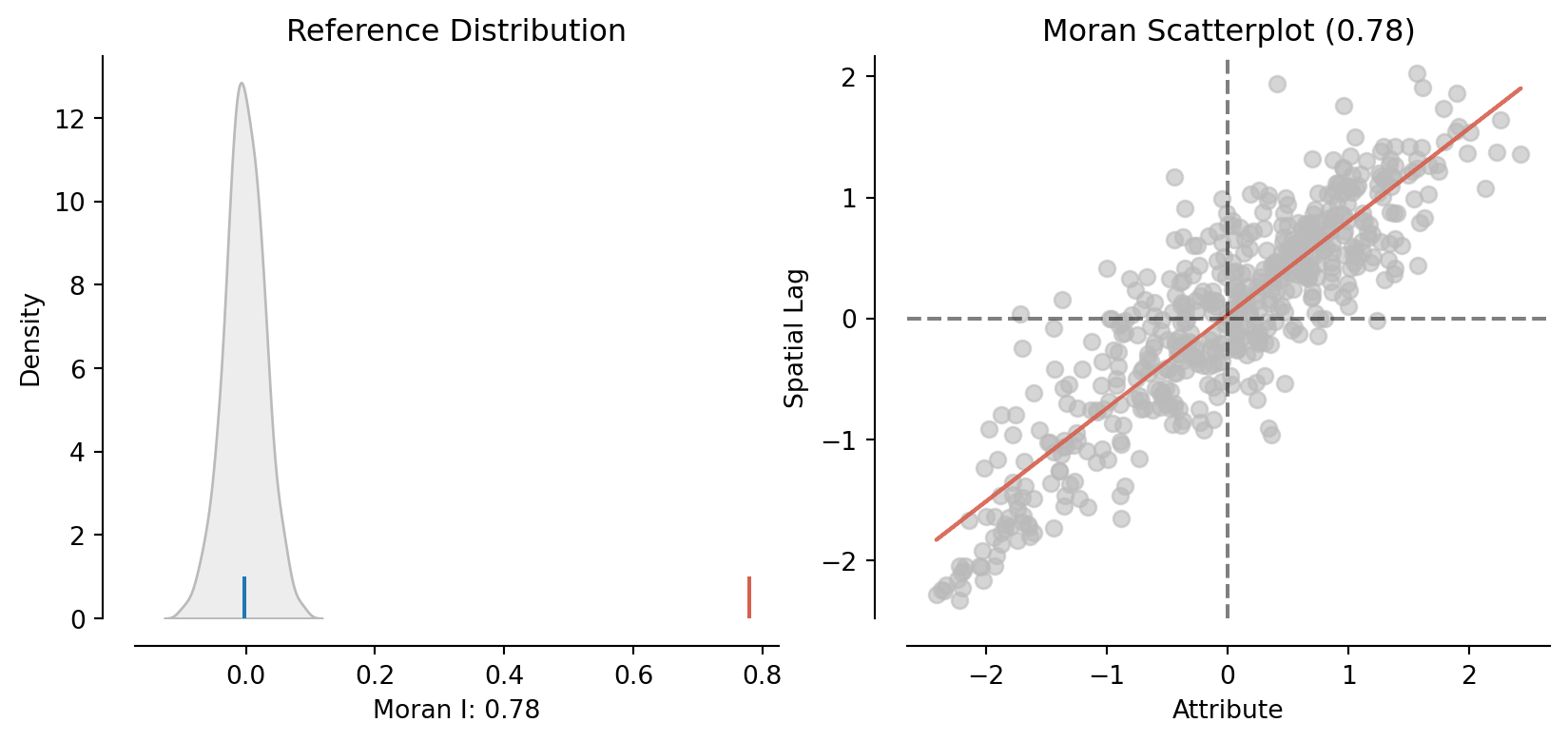
Practice
- Plot a chloropleth map of
D9_diff(9-th decile income) (castD9as a float first) - Compute its spatially lagged value
- Plot
D9and its spatially lagged value: what is the relationship between the two? - Compute Moran’s I
moran = esda.moran.Moran(constituencies["D9_diff"].astype(float), W)
plot_moran(moran)(<Figure size 960x384 with 2 Axes>,
array([<Axes: title={'center': 'Reference Distribution'}, xlabel='Moran I: 0.75', ylabel='Density'>,
<Axes: title={'center': 'Moran Scatterplot (0.75)'}, xlabel='Attribute', ylabel='Spatial Lag'>],
dtype=object))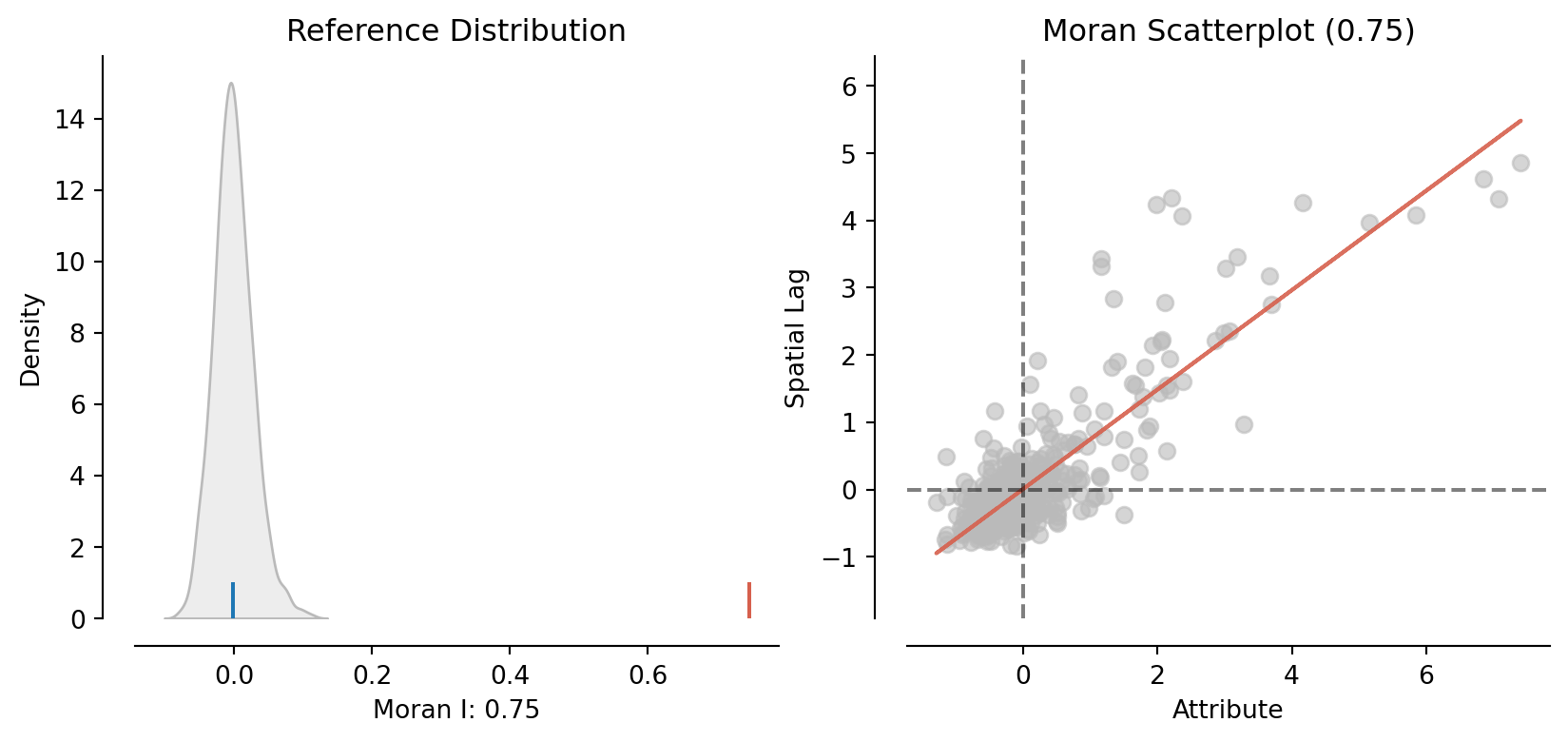
constituencies.plot("D9_diff", cmap="Reds")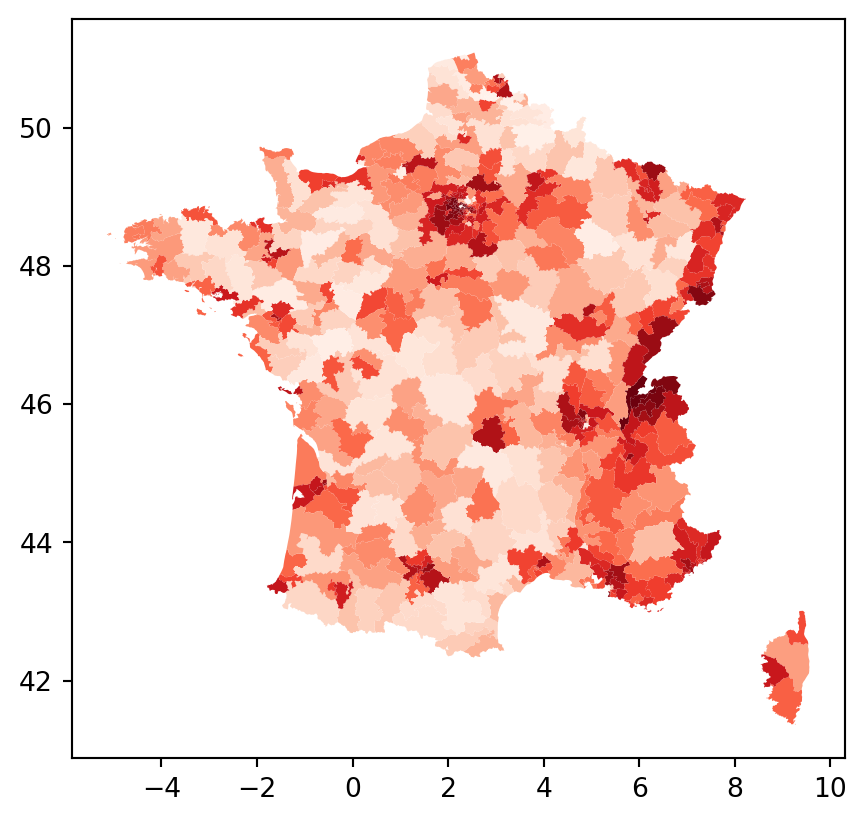
Local autocorrelation
One Local Indicator of Spatial Association (LISA) is local Moran’s I: global Moran’s I, expect we don’t sum over the i’s
\(I_i = \frac{n}{\sum_{i} z_{i}^2} z_{i} \sum_{j} w_{i,j}z_{j}\)
lisa = esda.moran.Moran_Local(constituencies["vote_share_rn"], W)
lisa.Is/opt/python/lib/python3.13/site-packages/esda/moran.py:1350: RuntimeWarning: invalid value encountered in divide
self.z_sim = (self.Is - self.EI_sim) / self.seI_simarray([ 5.79424752e-01, 9.88925307e-01, 3.48507625e-01, 9.35026207e-02,
3.12888167e-01, 8.24897842e-01, -1.25263747e-01, 9.06920602e-01,
9.26783838e-01, 1.38754803e+00, 1.40285754e+00, -1.05035562e-02,
8.19739723e-01, 6.11477304e-02, 2.77357751e-02, 1.79380083e+00,
7.42958401e-02, 7.13910752e-01, -1.98169549e-02, 1.37322264e-01,
1.63967005e-01, -2.47265333e-02, -0.00000000e+00, 3.23873650e-01,
2.48517940e+00, 2.75701188e+00, 1.70449698e+00, -1.58206529e-01,
-8.14702695e-02, 1.18986244e-01, 2.53174694e-01, -1.02672730e-01,
-1.54155086e-02, 2.58171656e-02, 2.68546977e-02, 3.43235069e-01,
2.47691054e-01, 2.96179373e-01, 1.36454129e+00, 1.44775693e+00,
9.42053418e-02, 4.02394253e-02, 1.97047060e+00, 1.13667614e-01,
2.52146643e+00, 3.56473156e+00, 1.14930115e+00, 1.17335681e+00,
3.13413875e+00, 2.83922544e+00, 9.60659579e-02, 2.58408250e+00,
5.30615784e+00, 5.48890188e+00, 3.65903257e-01, 4.55052128e+00,
4.87068166e+00, 3.74737822e+00, 5.16031545e+00, 4.19224185e+00,
4.62637617e+00, 4.52929475e+00, 1.59978600e+00, 1.17804244e+00,
2.91669832e+00, 5.38406562e-01, 5.00634563e-01, 3.15589980e+00,
1.69755895e+00, 8.55677834e-01, 5.93958713e-01, 1.76636973e+00,
3.29483049e+00, 3.02526901e+00, 3.93050870e+00, -5.20289444e-01,
2.25870942e+00, 1.08669322e+00, -4.06522442e-02, 3.09635879e+00,
3.29975029e-02, 1.70319198e-01, 1.57884791e+00, 1.18592404e+00,
3.45042594e-01, 2.51850089e-01, 7.01140706e-01, 1.27575648e-02,
4.56957189e-01, 8.21536916e-01, 2.18248678e-03, 3.42894955e-01,
1.68311913e+00, 2.39812304e+00, 1.49995325e+00, -4.02316305e-02,
2.22271167e+00, 9.89781427e-01, 2.90597784e-01, 1.48844443e+00,
2.06554395e-01, 2.58297934e-01, -3.04278903e-01, 9.05335362e-01,
-1.62553252e-01, 1.30026941e+00, -4.15726652e-01, -2.29787771e-01,
7.00528600e-01, 1.10907471e-01, 2.73189899e+00, 3.25895727e+00,
1.45217883e+00, 2.49283314e+00, 2.81186968e+00, 3.30151349e+00,
1.74288049e+00, 2.11219195e-01, 1.14398828e+00, 8.28083086e-01,
9.40890363e-01, 1.44428172e-01, -9.57537177e-02, 7.79064501e-01,
5.11745666e-01, -7.69226002e-02, 3.16159333e-01, 2.94033218e-02,
9.67503051e-01, 5.05823108e-01, 2.83858330e-01, 5.29909648e-01,
-2.06094261e-02, 1.82628202e+00, -4.24077887e-02, -6.55306653e-02,
8.50001656e-01, 1.33244086e+00, 1.18647672e+00, 2.17784018e-01,
-6.12800832e-02, -1.46183761e-01, 2.97139022e+00, 3.15803899e+00,
2.84935962e+00, 4.09514123e-01, 5.10286548e-01, 1.93541422e+00,
1.98553165e-01, 4.20891368e-02, 5.45127823e-02, 1.92332055e-01,
1.21613170e-01, 1.05657155e-01, 4.20208673e-01, 2.81538142e-01,
5.12524981e-01, -2.46203720e-02, 4.15022366e-01, -1.08526558e-01,
3.06288642e-01, 3.92813147e-01, 4.78513586e-01, 6.90631438e-02,
-4.79518589e-02, 3.60153107e-01, 1.82705316e-01, 3.61782559e-01,
6.15019900e-01, 3.70142998e-01, 9.23386958e-02, 9.11971151e-01,
8.70520843e-01, -3.24646581e-01, 4.73437257e-01, 1.46883204e+00,
2.61843871e+00, 2.10688366e-01, -8.26927295e-02, 1.47534520e-02,
4.56830275e-02, 1.04303287e-01, -1.82945744e-02, 6.31108292e-02,
-4.68355690e-02, -1.33216478e-02, 1.77183392e+00, 1.74745389e+00,
1.29032517e+00, -4.28227904e-03, -1.82165813e-02, 1.91001878e-01,
3.97848456e-01, 3.08249939e+00, -1.45798882e-01, 6.11845203e-01,
4.50029155e-01, 9.09703204e-02, 2.84924795e-01, 8.03747041e-02,
3.20169942e+00, 2.12213150e+00, 7.85002532e-01, 1.35700462e+00,
1.89441254e-01, 1.48867165e+00, 6.70957869e-01, 1.24973197e+00,
1.60965093e-02, 3.42593815e-01, -2.61759140e-01, 1.34615420e+00,
9.91455859e-01, 1.02690628e+00, 5.48627965e-01, 4.42149186e-02,
-5.32605896e-03, 2.94146049e+00, 2.10785040e+00, 1.07367973e+00,
5.49774940e-01, 6.54111344e-01, 4.17327226e-01, 3.63750394e-02,
1.79106348e+00, 1.38454125e-01, 5.12039567e-01, 2.24170593e-02,
4.70411928e-01, 1.79617381e+00, 2.74923609e+00, -1.69456124e-01,
2.06975639e-03, 5.11287271e-01, 0.00000000e+00, 4.00348526e-01,
7.60382047e-01, 9.16355674e-01, 3.02051483e-01, 1.49239988e+00,
2.64043159e+00, -1.08398642e-01, -2.29259962e-02, 6.91443225e-01,
8.94204180e-01, 5.96233170e-02, 4.39191849e-04, 2.09330337e+00,
8.62538231e-01, 7.52011765e-01, 7.60384441e-01, 2.32662712e+00,
1.30730050e+00, 1.74965880e+00, 3.07252352e+00, 5.78889488e-01,
1.19555329e-01, 2.31697555e-01, 1.75435856e-02, -2.40264396e-02,
8.69690967e-02, 3.95275720e-01, -2.77904546e-02, 7.04733298e-02,
0.00000000e+00, -1.08617828e-01, 5.03257817e-01, 3.07911405e+00,
7.49271292e-02, 0.00000000e+00, 1.50366792e+00, 3.49201588e+00,
5.01264555e-01, 1.20186471e-01, 1.82390683e+00, 3.58278633e-01,
8.31512038e-01, -4.10591594e-02, 1.12098582e+00, 1.21503582e+00,
-2.61845309e-02, 2.40728786e-01, 8.53187774e-02, 3.17734797e+00,
5.65084490e-02, 6.42112323e-02, 1.95820083e-01, 6.29224598e-01,
3.09752510e-01, 3.90199438e+00, 5.11510030e+00, 4.43362601e+00,
4.81736065e+00, 5.25387720e+00, 3.47789569e+00, 4.17980233e+00,
4.35436558e+00, 9.75683211e-02, 2.84194535e-01, 1.85856610e+00,
3.04383619e+00, 2.29180748e+00, 1.51425873e+00, 1.96138499e+00,
9.18175801e-01, 3.70361284e+00, 2.20047080e+00, -3.44602925e-01,
3.51415175e+00, 1.25683251e+00, 2.12801095e+00, 1.48043114e+00,
8.38722592e-02, 2.18100265e-01, 1.83910413e-02, 4.82253023e-01,
4.49639564e-01, 2.18792083e-01, -6.10555829e-02, 8.42442858e-03,
4.78527898e-01, 7.98747751e-01, 2.33055823e-02, 3.45017707e-01,
1.70883960e-01, -1.51897743e-02, -0.00000000e+00, 2.19816571e-01,
1.13635832e+00, 2.05987422e+00, 1.13695150e-02, 3.62864992e-02,
5.47745892e-01, 4.74759047e-01, 4.10741648e-01, -3.01369000e-02,
-8.09462004e-03, 6.13663897e-01, -4.94498333e-02, 7.68270774e-01,
-1.15930142e-01, 1.07157130e+00, 3.18578686e-01, 2.01039048e-02,
1.96887363e-01, 3.33692194e-01, 1.43430061e+00, 1.02137669e+00,
1.51967396e+00, 6.12561440e-01, 4.57202396e-01, 1.02198893e-02,
8.32518451e-02, 9.17135242e-01, 2.38647724e+00, 5.54203393e-02,
4.35788239e-02, 6.58410559e-02, 8.47811802e-01, 5.44884396e-01,
7.05546329e-01, 2.86429117e-01, -4.27529600e-02, 1.17828835e+00,
2.11222866e-01, 1.53207322e-01, -1.16329928e-02, 5.20705470e-03,
2.86304601e-01, 1.83217718e-01, 2.76379832e-02, -5.26464959e-02,
-2.07755015e-02, 7.91113078e-02, 4.85859488e-01, 1.21554960e-01,
3.21105300e-02, 5.05050623e-02, 1.94524383e-01, 2.22110235e-01,
-2.47814451e-01, -2.54067786e-02, 2.16612670e-01, 2.91660760e-01,
-3.03985600e-02, 4.25433990e-01, 1.30570010e-01, -1.68536243e-02,
3.76561615e-02, 1.83803519e-02, 1.23295473e-02, -1.17269388e-02,
4.59173805e-03, -2.14560109e-02, 1.59544306e-02, 4.13152118e-03,
1.01189647e-03, 6.15413191e-01, 2.74615128e-01, 1.70053419e+00,
2.37449605e-02, -4.02985956e-02, 1.74369137e+00, 5.03138732e-01,
1.58639175e+00, -1.50879442e-02, -5.15752311e-04, 5.30372146e-03,
-1.16813444e-01, 6.13136914e-01, 6.92836924e-01, 5.73806851e-01,
-1.06127496e-01, 1.34865636e-01, 2.69740094e+00, 2.33479995e-01,
-9.22112576e-03, 1.26329075e-03, -2.52585528e-01, 2.52405101e-01,
2.14718661e-03, 3.54029743e-01, 6.19330983e-02, 8.24750244e-01,
2.62205341e-01, 3.63996200e-01, 2.63784660e-01, 8.12508261e-01,
1.55333662e-01, 6.14627535e-01, 3.75097917e-01, 2.73676585e-01,
1.08440210e+00, 1.89722181e-01, -6.10932741e-02, 2.53458154e-02,
4.19547823e-02, 1.98813344e+00, 8.82451051e-01, 5.07341619e-01,
6.02967390e-02, 8.30283428e-01, 1.20173172e-01, 2.69457937e-01,
-3.98251622e-02, 1.29369744e-02, 6.01611091e-01, -2.12235934e-02,
9.97359636e-02, 3.44458059e-02, -2.09903378e-01, 1.70403964e+00,
1.36669557e+00, 7.10199153e-01, -3.50648693e-02, -1.76394409e-01,
9.67987453e-03, -2.87070713e-08, 2.28975795e-01, 1.84228683e-01,
2.34032954e-01, 2.45268843e-01, 2.84829380e-01, 4.84907331e-01,
1.55813743e-01, 7.64179470e-01, 3.09957578e-01, 1.28960924e+00,
7.96548779e-01, 1.57967088e+00, 3.25739812e-01, 4.98904341e-01,
3.73921757e-01, 3.06511387e-02, 1.21472305e+00, 1.55789710e+00,
6.60601726e-01, 4.64702169e-01, -1.49765826e-01, 8.67988573e-02,
1.99835938e-02, -2.73940159e-03, 3.46559423e-02, -1.72224335e-02,
5.48203157e-02, 3.37493794e-01, 9.90672759e-02, 6.45834912e-01,
7.73266648e-01, 4.64775424e-01, -1.13854918e-02, 3.60500213e-01,
1.90271251e-01, -7.57584033e-02, 2.91739444e-01, -4.85033794e-02,
3.41601264e-01, -9.27450200e-03, 1.97961112e+00, -4.46914851e-03,
1.77118107e-02, -9.86883804e-03, 1.16671751e-01, -4.92884640e-02,
9.94937072e-02, 1.27462575e+00, -3.62936487e-03, 1.24311111e-01,
2.47265519e-01, -1.04231823e-02, 2.82315051e+00, 6.14006753e-01,
9.42654177e-01, 1.17474992e+00, -1.89648058e-02, 1.21860624e-01,
3.67344069e-01, 2.14970044e-01, 1.19577932e+00, 1.28914042e+00,
1.49288061e+00, 9.14251674e-01, 1.79476656e-01, 3.29946767e-01,
9.48627308e-02, 8.55177993e-02, 2.30666555e-01, -5.30234222e-02,
1.46999038e-01, 4.58705007e-01, 3.52191601e-01, 3.41343557e-02,
-2.88194629e-01, 3.46731968e-01, -4.67076659e-02, 4.52700954e-01,
2.47949894e-01, 1.63147628e-02, -8.94117501e-03])plt.hist(lisa.Is)(array([165., 189., 68., 47., 17., 22., 13., 6., 7., 5.]),
array([-0.52028944, 0.08062969, 0.68154882, 1.28246795, 1.88338709,
2.48430622, 3.08522535, 3.68614448, 4.28706361, 4.88798275,
5.48890188]),
<BarContainer object of 10 artists>)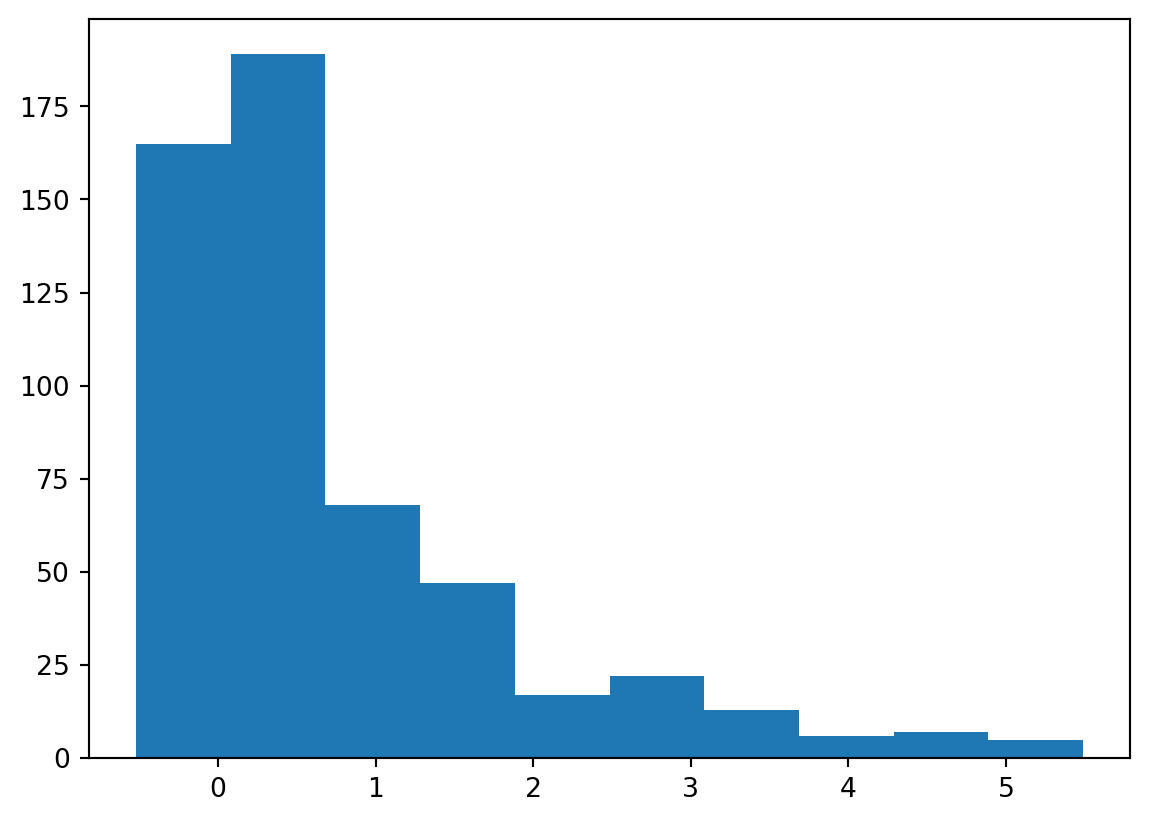
lisa.Is.mean()np.float64(0.7707928329691234)constituencies.assign(i=lisa.Is).plot(column="i", cmap = "Reds")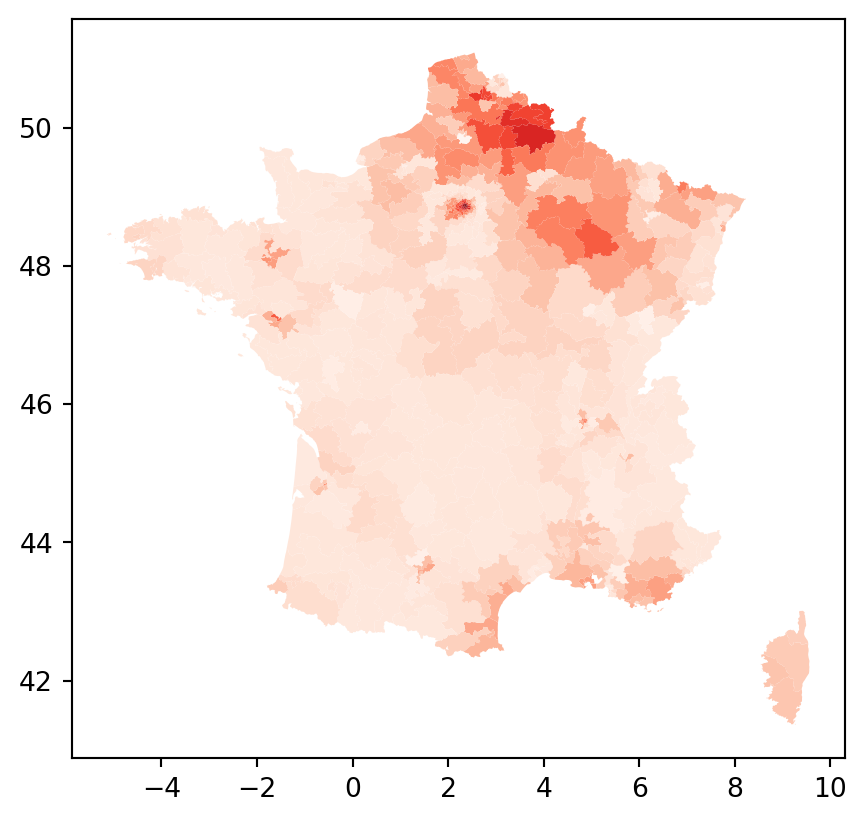
lisa_cluster(lisa, constituencies, p=1)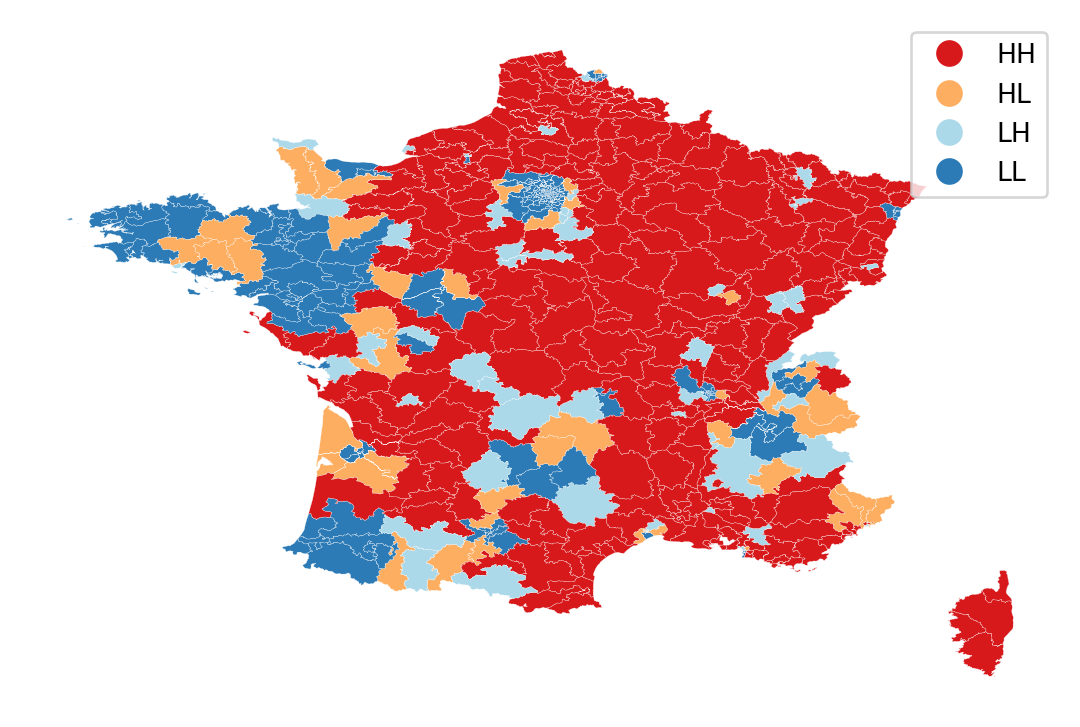
lisa_cluster(lisa, constituencies, p=0.05)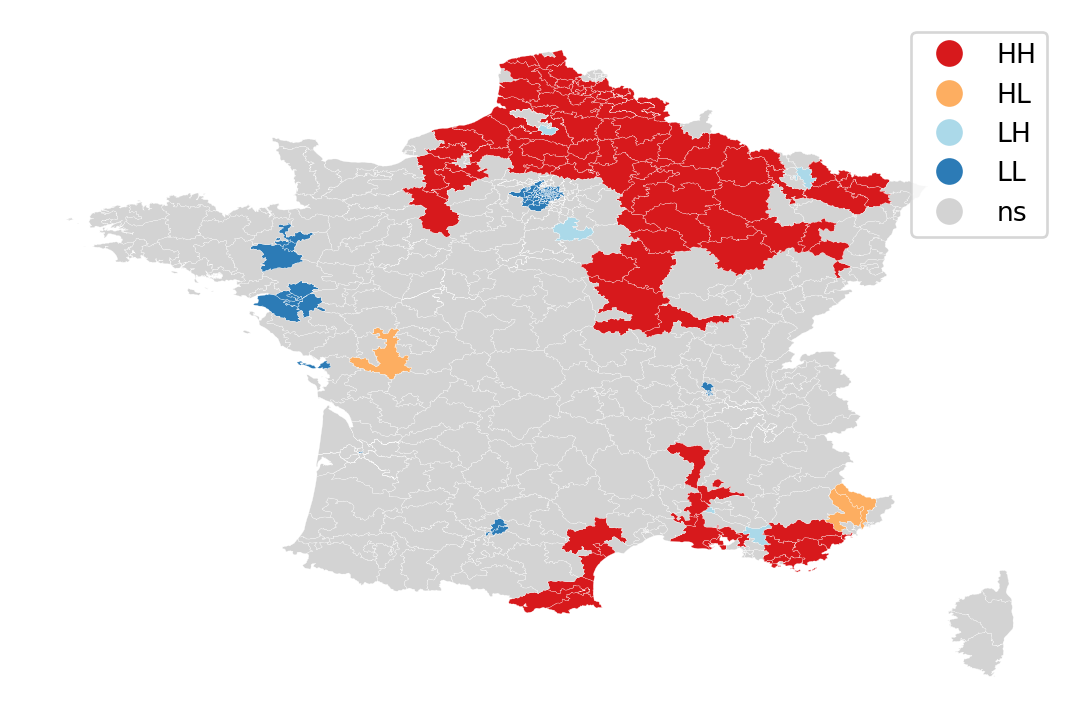
Practice
- Compute local Moran’s I for
D1_diff - Plot the Is, and the LL/HL/LH/HH clusters
Standard geographic regression models
- If the data generating process is explicitly spatial, we want to include geography in the analysis
- If some omitted variables is spatial, resulting errors will be spatial
Let’s see if we can predict/explain vote_share_rn with our explanatory variables
constituencies["D1_diff"] = constituencies["D1_diff"].astype(float)
constituencies["D9_diff"] = constituencies["D9_diff"].astype(float)
ols_model = spreg.OLS(
constituencies[['vote_share_rn']].values,
constituencies[['unemployement_rate', 'mean_age', 'D1_diff', 'D9_diff']].values,
# Dependent variable name
name_y="vote_share_rn",
# Independent variable name
name_x=['unemployement_rate', 'mean_age', 'D1_diff', 'D9_diff'],
)print(ols_model.summary)REGRESSION RESULTS
------------------
SUMMARY OF OUTPUT: ORDINARY LEAST SQUARES
------------------------------------------------------------------------------------
Data set : unknown
Weights matrix : None
Dependent Variable :vote_share_rn Number of Observations: 539
Mean dependent var : 31.6400 Number of Variables : 5
S.D. dependent var : 10.7351 Degrees of Freedom : 534
R-squared : 0.4494
Adjusted R-squared : 0.4452
Sum squared residual: 34140.1 F-statistic : 108.9422
Sigma-square : 63.933 Prob(F-statistic) : 7.867e-68
S.E. of regression : 7.996 Log likelihood : -1882.832
Sigma-square ML : 63.340 Akaike info criterion : 3775.664
S.E of regression ML: 7.9586 Schwarz criterion : 3797.113
------------------------------------------------------------------------------------
Variable Coefficient Std.Error t-Statistic Probability
------------------------------------------------------------------------------------
CONSTANT -59.14834 11.61176 -5.09383 0.00000
unemployement_rate 3.02469 0.53091 5.69717 0.00000
mean_age 1.36535 0.12838 10.63541 0.00000
D1_diff 0.00313 0.00050 6.22522 0.00000
D9_diff -0.00054 0.00004 -14.03249 0.00000
------------------------------------------------------------------------------------
REGRESSION DIAGNOSTICS
MULTICOLLINEARITY CONDITION NUMBER 86.220
TEST ON NORMALITY OF ERRORS
TEST DF VALUE PROB
Jarque-Bera 2 7.609 0.0223
DIAGNOSTICS FOR HETEROSKEDASTICITY
RANDOM COEFFICIENTS
TEST DF VALUE PROB
Breusch-Pagan test 4 47.549 0.0000
Koenker-Bassett test 4 64.950 0.0000
================================ END OF REPORT =====================================constituencies.assign(predy=ols_model.predy).plot(column="predy", cmap = "Reds")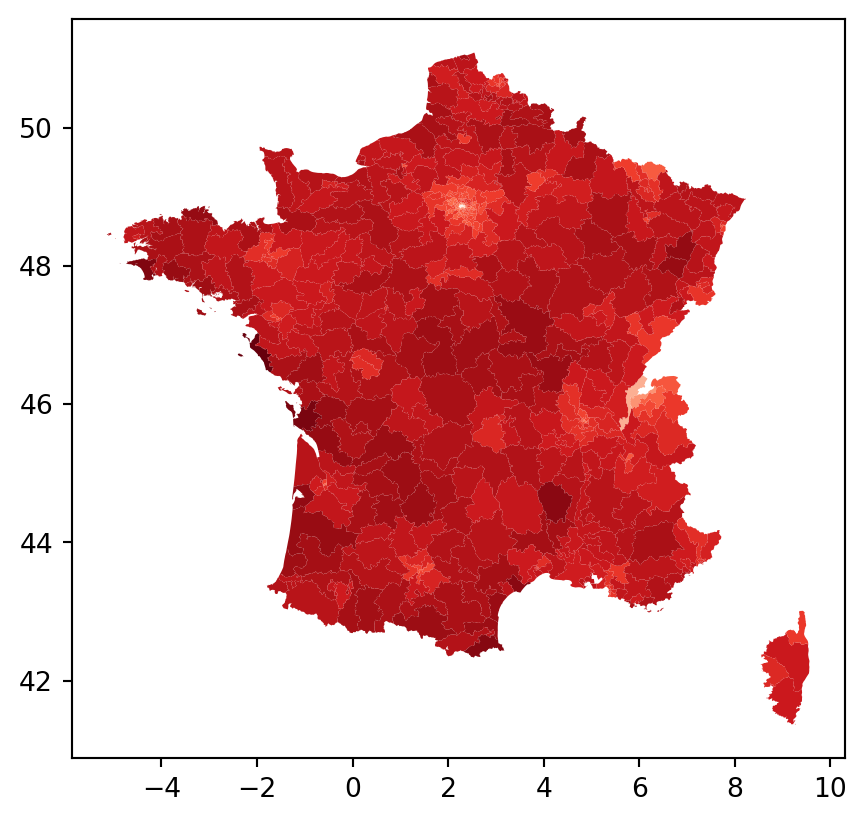
constituencies.assign(u=ols_model.u).plot(column="u", cmap = "seismic", legend = True)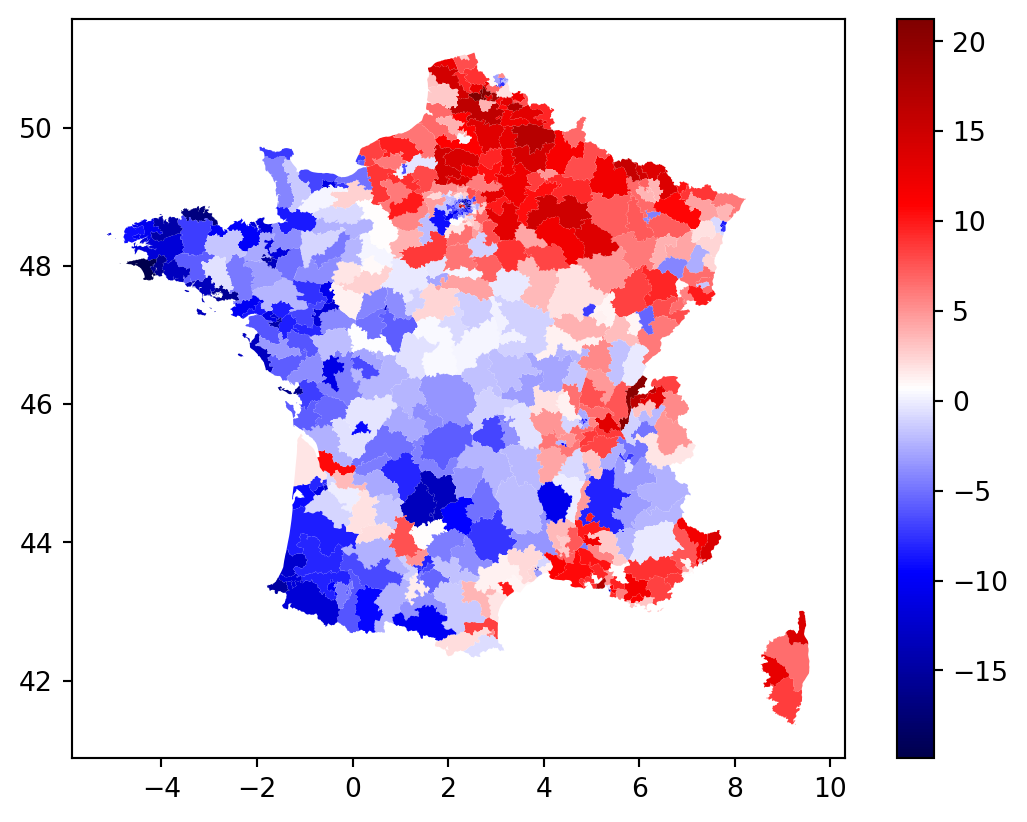
constituencies['u'] = ols_model.u
# constituencies.boxplot(column = ['residuals'], by = "dep", figsize = (20, 5))
constituencies["lagged_u"] = weights.lag_spatial(W, constituencies["u"])
fig, ax = plt.subplots(figsize=(9, 9))
ax.scatter(constituencies["u"], constituencies["lagged_u"])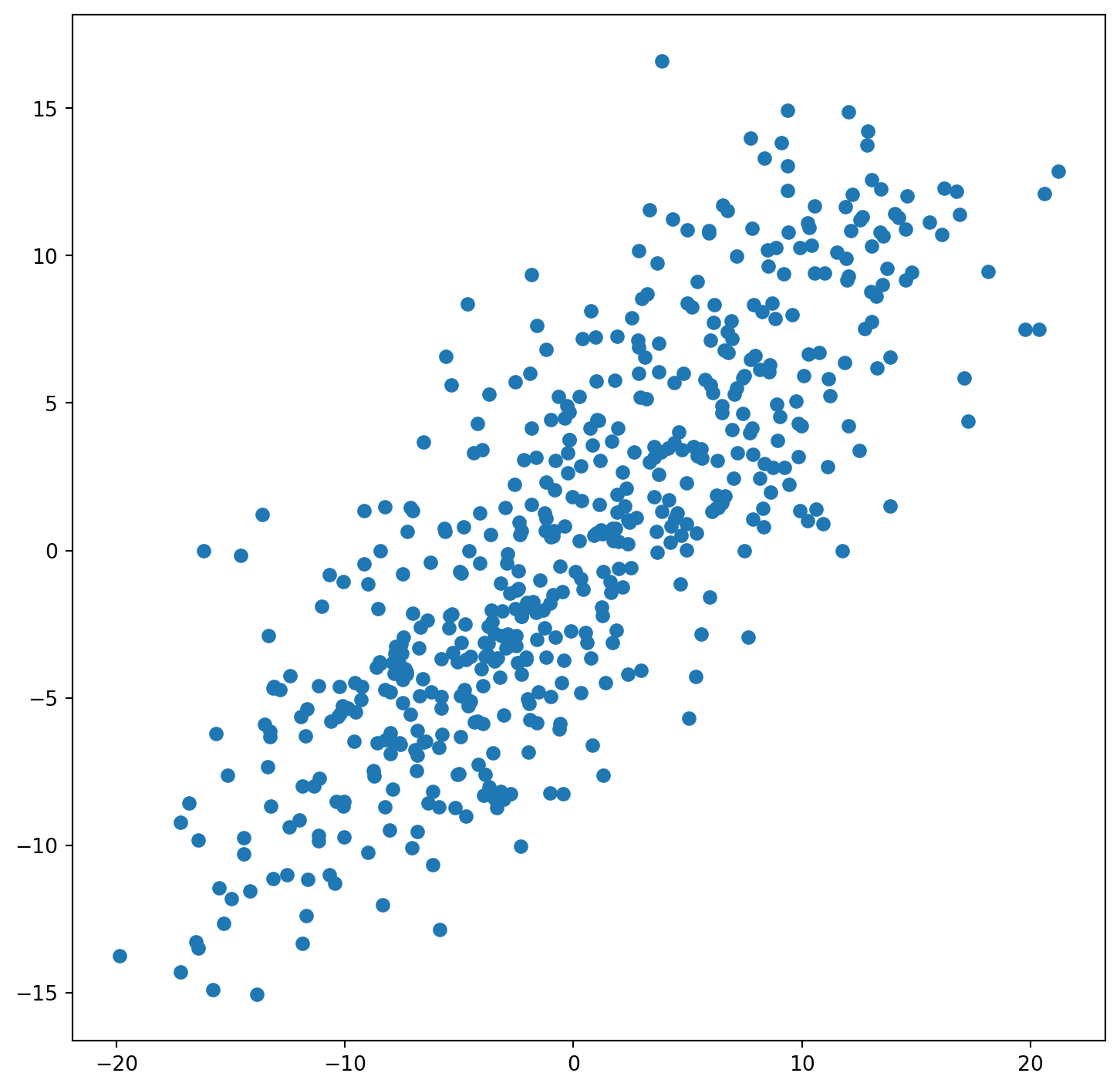
SLX model
Add spatially lagged values as exogenous variables, i.e. \(WX_j\) influence \(Y_i\)
constituencies["lagged_unemployement_rate"] = weights.lag_spatial(W, constituencies["unemployement_rate"])
constituencies["lagged_mean_age"] = weights.lag_spatial(W, constituencies["mean_age"])
constituencies["lagged_D1_diff"] = weights.lag_spatial(W, constituencies["D1_diff"])
constituencies["lagged_D9_diff"] = weights.lag_spatial(W, constituencies["D9_diff"])
variables = ['unemployement_rate', 'mean_age', 'D1_diff', 'D9_diff']
lagged_variables = ["lagged_" + x for x in variables]
all_variables = variables + lagged_variables
slx_model = spreg.OLS(
constituencies[['vote_share_rn']].values,
constituencies[all_variables].values,
# Dependent variable name
name_y="vote_share_rn",
# Independent variable name
name_x=all_variables
)
print(slx_model.summary)REGRESSION RESULTS
------------------
SUMMARY OF OUTPUT: ORDINARY LEAST SQUARES
------------------------------------------------------------------------------------
Data set : unknown
Weights matrix : None
Dependent Variable :vote_share_rn Number of Observations: 539
Mean dependent var : 31.6400 Number of Variables : 9
S.D. dependent var : 10.7351 Degrees of Freedom : 530
R-squared : 0.4732
Adjusted R-squared : 0.4653
Sum squared residual: 32661.4 F-statistic : 59.5100
Sigma-square : 61.625 Prob(F-statistic) : 5.854e-69
S.E. of regression : 7.850 Log likelihood : -1870.899
Sigma-square ML : 60.596 Akaike info criterion : 3759.798
S.E of regression ML: 7.7844 Schwarz criterion : 3798.406
------------------------------------------------------------------------------------
Variable Coefficient Std.Error t-Statistic Probability
------------------------------------------------------------------------------------
CONSTANT -59.35288 11.79147 -5.03354 0.00000
unemployement_rate 2.16722 0.58391 3.71157 0.00023
mean_age 1.30285 0.19924 6.53918 0.00000
D1_diff 0.00310 0.00062 5.02964 0.00000
D9_diff -0.00046 0.00007 -6.09763 0.00000
lagged_unemployement_rate 1.74391 0.50921 3.42472 0.00066
lagged_mean_age 0.05791 0.19780 0.29279 0.76980
lagged_D1_diff -0.00026 0.00060 -0.44086 0.65949
lagged_D9_diff -0.00011 0.00009 -1.33309 0.18307
------------------------------------------------------------------------------------
REGRESSION DIAGNOSTICS
MULTICOLLINEARITY CONDITION NUMBER 122.403
TEST ON NORMALITY OF ERRORS
TEST DF VALUE PROB
Jarque-Bera 2 3.904 0.1420
DIAGNOSTICS FOR HETEROSKEDASTICITY
RANDOM COEFFICIENTS
TEST DF VALUE PROB
Breusch-Pagan test 8 83.573 0.0000
Koenker-Bassett test 8 104.473 0.0000
================================ END OF REPORT =====================================slx_model_2 = spreg.OLS(
constituencies[['vote_share_rn']].values,
constituencies[variables].values,
name_y="vote_share_rn",
name_x=variables,
w = W,
slx_lags = 1
)
print(slx_model_2.summary)REGRESSION RESULTS
------------------
SUMMARY OF OUTPUT: ORDINARY LEAST SQUARES WITH SPATIALLY LAGGED X (SLX)
------------------------------------------------------------------------------------
Data set : unknown
Weights matrix : unknown
Dependent Variable :vote_share_rn Number of Observations: 539
Mean dependent var : 31.6400 Number of Variables : 9
S.D. dependent var : 10.7351 Degrees of Freedom : 530
R-squared : 0.4732
Adjusted R-squared : 0.4653
Sum squared residual: 32661.4 F-statistic : 59.5100
Sigma-square : 61.625 Prob(F-statistic) : 5.854e-69
S.E. of regression : 7.850 Log likelihood : -1870.899
Sigma-square ML : 60.596 Akaike info criterion : 3759.798
S.E of regression ML: 7.7844 Schwarz criterion : 3798.406
------------------------------------------------------------------------------------
Variable Coefficient Std.Error t-Statistic Probability
------------------------------------------------------------------------------------
CONSTANT -59.35288 11.79147 -5.03354 0.00000
unemployement_rate 2.16722 0.58391 3.71157 0.00023
mean_age 1.30285 0.19924 6.53918 0.00000
D1_diff 0.00310 0.00062 5.02964 0.00000
D9_diff -0.00046 0.00007 -6.09763 0.00000
W_unemployement_rate 1.74391 0.50921 3.42472 0.00066
W_mean_age 0.05791 0.19780 0.29279 0.76980
W_D1_diff -0.00026 0.00060 -0.44086 0.65949
W_D9_diff -0.00011 0.00009 -1.33309 0.18307
------------------------------------------------------------------------------------
REGRESSION DIAGNOSTICS
MULTICOLLINEARITY CONDITION NUMBER 122.403
TEST ON NORMALITY OF ERRORS
TEST DF VALUE PROB
Jarque-Bera 2 3.904 0.1420
DIAGNOSTICS FOR HETEROSKEDASTICITY
RANDOM COEFFICIENTS
TEST DF VALUE PROB
Breusch-Pagan test 8 83.573 0.0000
Koenker-Bassett test 8 104.473 0.0000
================================ END OF REPORT =====================================Beware of marginal effect computation :
- effect of increasing \(x_i\)
- effect of increasing all \(x_j\) on \(y_i\)
- effect of increasing \(x_i\) and \(x_j\) on \(y_i\)
Spatial Error model (SEM)
Allow for spatial autocorrelation in errors : \(Wu_j\) influence \(u_i\) (and thus \(Y_i\))
\(u_i = \lambda Wu_j + \epsilon_{i}\)
This imply heteroskedasticity hence OLS is not efficient
sem_model = spreg.GM_Error_Het(
constituencies[['vote_share_rn']].values,
constituencies[variables].values,
name_y="vote_share_rn",
name_x=variables,
w = W,
)
print(sem_model.summary)GM_Error_Het
REGRESSION RESULTS
------------------
SUMMARY OF OUTPUT: GM SPATIALLY WEIGHTED LEAST SQUARES (HET)
------------------------------------------------------------------------------------
Data set : unknown
Weights matrix : unknown
Dependent Variable :vote_share_rn Number of Observations: 539
Mean dependent var : 31.6400 Number of Variables : 5
S.D. dependent var : 10.7351 Degrees of Freedom : 534
Pseudo R-squared : 0.4055
N. of iterations : 1 Step1c computed : No
------------------------------------------------------------------------------------
Variable Coefficient Std.Error z-Statistic Probability
------------------------------------------------------------------------------------
CONSTANT -7.98323 8.47178 -0.94233 0.34602
unemployement_rate 0.30196 0.38959 0.77507 0.43830
mean_age 0.83133 0.12537 6.63103 0.00000
D1_diff 0.00194 0.00043 4.48594 0.00001
D9_diff -0.00052 0.00007 -7.33055 0.00000
lambda 0.82500 0.02364 34.89286 0.00000
------------------------------------------------------------------------------------
================================ END OF REPORT =====================================Spatial Lag Model (or Spatial Autoregressive Model)
\(WY_j\) influences \(Y_i\)
Violates exogeneity conditions
slm_model = spreg.GM_Lag(
constituencies[['vote_share_rn']].values,
constituencies[variables].values,
name_y="vote_share_rn",
name_x=variables,
w = W,
)
print(slm_model.summary)GM_Lag
REGRESSION RESULTS
------------------
SUMMARY OF OUTPUT: SPATIAL TWO STAGE LEAST SQUARES
------------------------------------------------------------------------------------
Data set : unknown
Weights matrix : unknown
Dependent Variable :vote_share_rn Number of Observations: 539
Mean dependent var : 31.6400 Number of Variables : 6
S.D. dependent var : 10.7351 Degrees of Freedom : 533
Pseudo R-squared : 0.5887
Spatial Pseudo R-squared: 0.4566
------------------------------------------------------------------------------------
Variable Coefficient Std.Error z-Statistic Probability
------------------------------------------------------------------------------------
CONSTANT -50.10444 10.56509 -4.74245 0.00000
unemployement_rate 2.43502 0.50534 4.81855 0.00000
mean_age 1.13751 0.13736 8.28134 0.00000
D1_diff 0.00267 0.00046 5.75318 0.00000
D9_diff -0.00045 0.00004 -10.19554 0.00000
W_vote_share_rn 0.19117 0.06749 2.83249 0.00462
------------------------------------------------------------------------------------
Instrumented: W_vote_share_rn
Instruments: W_D1_diff, W_D9_diff, W_mean_age, W_unemployement_rate
DIAGNOSTICS FOR SPATIAL DEPENDENCE
TEST DF VALUE PROB
Anselin-Kelejian Test 1 33.244 0.0000
SPATIAL LAG MODEL IMPACTS
Impacts computed using the 'simple' method.
Variable Direct Indirect Total
unemployement_rate 2.4350 0.5755 3.0106
mean_age 1.1375 0.2689 1.4064
D1_diff 0.0027 0.0006 0.0033
D9_diff -0.0005 -0.0001 -0.0006
================================ END OF REPORT =====================================| Model | Formula | When to Use |
|---|---|---|
| OLS | \(Y = X\beta + \epsilon\) | Baseline, no spatial effects |
| SLX | \(Y = X\beta + WX\gamma + \epsilon\) | Neighbor characteristics affect outcome |
| SEM | \(Y = X\beta + u\), \(u = \lambda Wu + \epsilon\) | Spatial error correlation (omitted variables) |
| SAR/Lag | \(Y = \rho WY + X\beta + \epsilon\) | Outcomes depend on neighbor outcomes |
Where: - \(W\) = spatial weights matrix - \(WX\) = spatially lagged explanatory variables - \(WY\) = spatially lagged dependent variable - \(\lambda\) = spatial error parameter - \(\rho\) = spatial lag parameter